How do you find the molecular weight of a protein?

How do you find the molecular weight of a protein?
If the sequence of amino acids is known for any given protein, we can roughly estimate the molecular weight of the protein by multiplying the average molecular weight of amino acids with the total number of amino acids present in the protein. The average molecular weight of an amino acid is known to be 110 Daltons(Da).
What is the protein sequence database?
The Protein database is a collection of sequences from several sources, including translations from annotated coding regions in GenBank, RefSeq and TPA, as well as records from SwissProt, PIR, PRF, and PDB. Protein sequences are the fundamental determinants of biological structure and function.
How do you find the molecular weight of a protein from an amino acid?
The molecular weight (mw) of an oligopeptide or a protein can be determined by summation of the mw of its corresponding amino acid sequence.
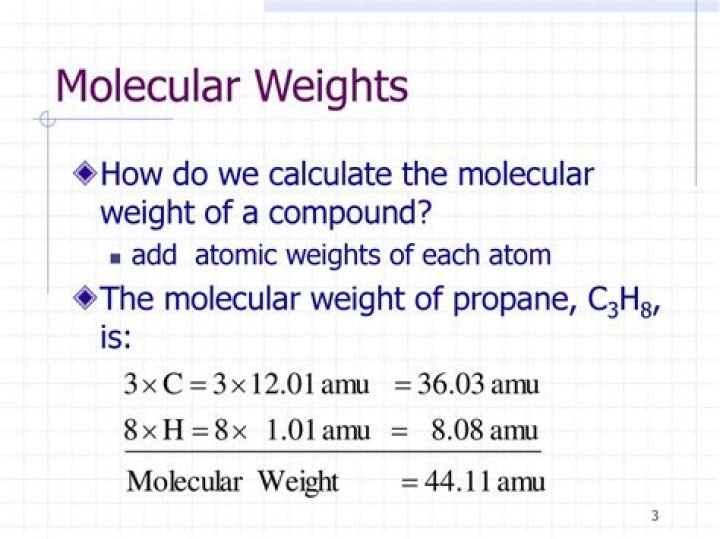
How do we calculate molecular weight?
molecular weight = (number of carbon atoms)(C atomic weight) + (number of H atoms)(H atomic weight) so we calculate as follows: molecular weight = (6 x 12.01) + (14 x 1.01)
How do you find the molecular weight of an unknown protein?
Use a graphing program, plot the log (MW) as a function of Rf. Generate the equation y = mx + b, and solve for y to determine the MW of the unknown protein. Run the standards and samples on an SDS-PAGE gel. Process the gel with the desired stain and then destain to visualize the protein bands.
How does NCBI determine protein structure?
How to: View the 3D structure of a protein
- Go to the Structure Home Page.
- Enter the PDB code in the search box and press the Go button.
- Click a structure image to access its record page.
- Scroll to the molecular graphic section and click on the spin icon to load an interactive view of the structure within the web page.
What is the secondary protein sequence database?
A protein secondary structure database (PSS) has been designed to correlate the Protein Sequence Database of the PIR-International with the atomic coordinates and bond connectivities database of the Protein Data Bank in the Brookhaven National Laboratory.
How do you find the molecular weight of a base pair of protein?
The molecular weight or molar mass of any double-stranded DNA fragment can therefore be calculated by multiplying its length (in bp) by 650 and the answer will be expressed as daltons or g/mol.
How do you find the molecular weight of CA hco3 2?
162.1146 g/mol
Calcium bicarbonate/Molar mass